Caris Assure Blood-based Profiling
Not available in all locations.
Caris Assure for Therapy Selection
Caris Assure® is a minimally invasive blood test that provides comprehensive molecular analysis of tumor biomarkers when tumor tissue is not feasible. The assay utilizes circulating Nucleic Acid Sequencing (cNAS), a novel approach to liquid biopsy that analyzes cell-free DNA and RNA from plasma, plus genomic DNA and messenger RNA from circulating white blood cells (WBCs), to distinguish:
- Somatic tumor variants – This is the most critical information needed by a physician to make treatment decisions about targeted therapies. By distinguishing somatic (acquired) variants from incidental CH or incidental germline results, Caris Assure provides confidence in therapy decisions and avoidance of off-target drugs.
- Incidental CH – Clonal hematopoiesis (CH) mutations from aging white blood cells may cause false positive results for therapy selection. Caris Assure distinguishes CH from somatic tumor mutations to improve accuracy. Read the 2025 Study >
- Incidental germline† – Caris Assure analyzes genomic DNA from circulating white blood cells and can distinguish incidental germline mutations from somatic mutations. A germline mutation is present in even non-tumor cells and may not be a target for therapy.
- Pharmacogenomics (DPYD) – Caris Assure directly analyzes plasma and white blood cells to identify certain inherited gene variants, including DPYD, that affect how chemotherapy medicines are metabolized. This comprehensive approach helps physicians reduce the risk of severe, life-threatening toxicity and personalize treatment safely.
Caris Assure has the potential to pick up more mutations, with a sensitivity of 93.8%* when compared to matched tissue collected within 30 days and using ≥5 ng of input material. Test results include genomic signatures, including microsatellite instability (MSI), blood tumor mutational burden (bTMB) and predicted HLA genotype (confirmatory testing required), providing deep molecular insights to help inform therapy decisions.
*See Technical Information for performance specifications
SPECIMEN TYPE(S)
Whole Blood
APPLICATION
Profiling for therapy selection
Less Invasive, More Intelligent Than Ever

TECHNOLOGY
Circulating Nucleic Acid
Sequencing (cNAS)
APPLICATION
Biomarker Analysis (including resistance mutations)
BIOLOGICAL COVERAGE
Plasma: cfDNA, cfRNA
White Blood Cells: gDNA, mRNA
VARIANT COVERAGE (PATHOGENIC AND LIKELY PATHOGENIC)
Tumor-Derived
Incidental CH
Incidental Germline†
GENES & DEPTH
23,000+ 10,000x (raw average for clinically relevant genes)
NEXT GENERATION SEQUENCING
Whole Exome
Whole Transcriptome
ALTERATIONS
SNV InDel CNA Fusions
GENOMIC SIGNATURES / OTHER
bTMB Predicted HLA Genotype MSI
SAMPLE QUANTITY
Two Tubes Whole Blood
PERFORMANCE IN ADVANCED/
METASTATIC PATIENTS
Compared to matched tissue collected within 30 days; based on ≥5 ng of cNAS input. Minimum reportable allele frequency is 0.1%.
Clinically Actionable SNV and InDel:
Sensitivity 93.8%
Specificity >99.9%
PPV 96.8%
Incidental Germline† :
Sensitivity >99%
Specificity >99%
PPV >99%
†Not a replacement for comprehensive germline testing. Incidental pathogenic alterations are reported, including ACMG recognized cancer genes. Negative results do not imply the patient does not harbor a germline mutation.
Caris Assure™ is intended for patients with previously diagnosed solid malignant neoplasms when tissue is not feasible and is to be used by qualified healthcare professionals. RNA results are intended for investigational purposes only. Not available in all locations.
Treat the Tumor, Not the False Positives
Caris Assure distinguishes somatic tumor variants from incidental clonal hematopoiesis (CH) and incidental germline results, helping physicians to select therapies that target the tumor.

Somatic Tumor
By distinguishing somatic (acquired) variants from incidental CH or incidental germline results, Caris Assure provides confidence in therapy decisions and avoidance of off-target drugs.

Incidental Germline
Caris Assure analyzes genomic DNA from circulating white blood cells found in the buffy coat and can distinguish incidental germline† mutations from somatic mutations.
Order Profiling
Email the completed form(s) to CustomerSupport@CarisLS.com, or fax to 1.866.479.4925. When specimen is being prepared for shipment, please include completed forms with the shipper. Not available in all locations.
Tour Our Blood Lab
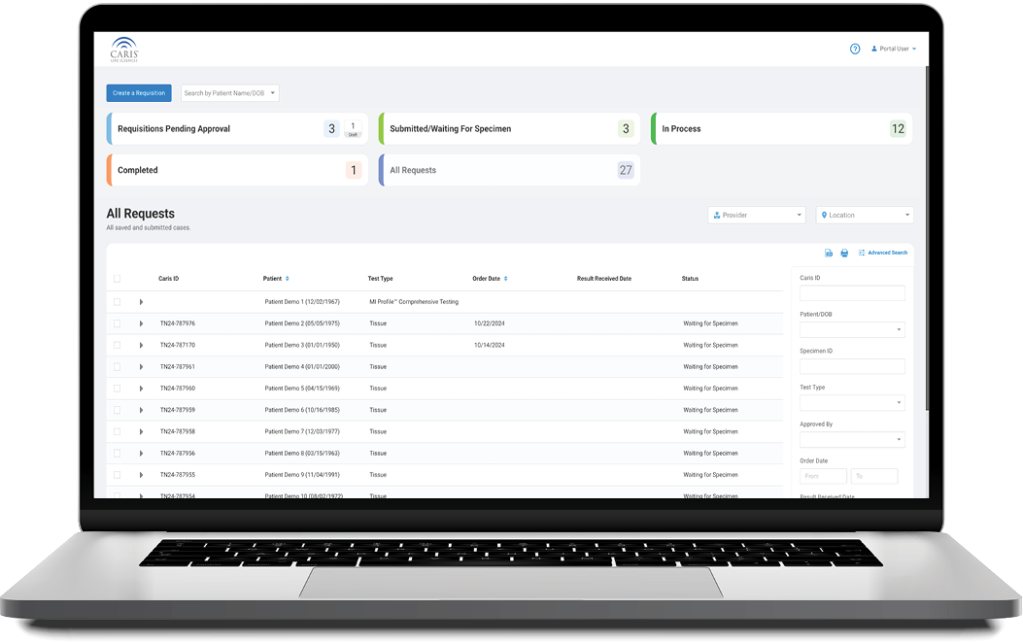
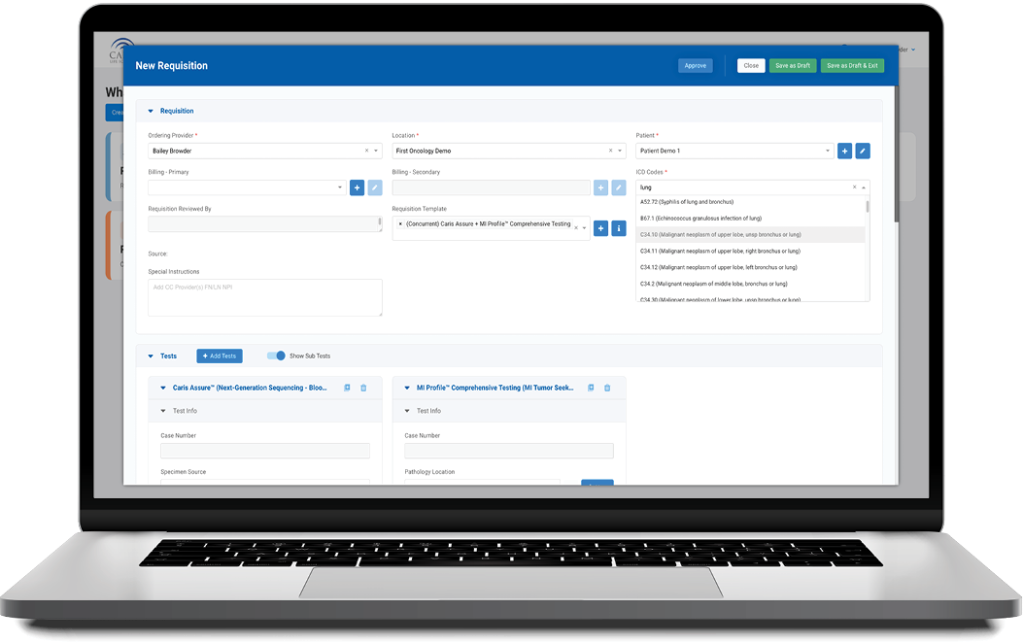
Caris+Portal
Convenient Access to Caris Profiling
Caris+™Portal provides easy access for users to electronically submit orders, track case progress, view results and review Caris Life Sciences’ profiling information in one convenient location.
New users will select the Register link at the bottom of the login page and enter their name and their clinic or institution email address to verify the account.

Complete Molecular Intelligence® Report
The Caris Molecular Profiling Report delivers high impact results, including potentially relevant, actionable clinical information, in an easy-to-interpret format. Every report includes access to the MI Portal and the Clinical Trials Connector™, which matches each patient’s unique biomarker expression profile to open, pertinent and clinical trial opportunities.
| Sample Requirements
(see requisition for full details) | 2 x 10 mL tubes whole blood | |
| Number of Genes | 23,000+ genes | |
| Average Depth of Coverage (DNA) | 10,000x (raw average for clinically relevant genes) | |
| Sensitivity | 93.8% for clinical genes in advanced/metastatic cases >99% for incidental germline | |
| Specificity | >99.9% for SNV and InDel >99% for Incidental Germline | |
| Alterations | SNV, InDel, CNA, Fusions | |
| Genomic Signatures/Other | Blood Tumor Mutational Burden (bTMB) Predicted Human Leukocyte Antigen (HLA) Genotype Microsatellite Instability (MSI) | |

I wanted to know everything I could about my cancer. Molecular profiling helped create the best, most personalized treatment plan for my survival.”
Patient, Shawna Stengle, Triple Negative Breast Cancer Survivor
Patient, Shawna Stengle, Triple Negative Breast Cancer Survivor
Discover
More
Caris MI Profile® comprehensive testing delivers whole exome sequencing (WES – DNA) and whole transcriptome sequencing (WTS – RNA) for 23,000+ genes, as well as protein analysis and AI-predictive algorithms.
Caris partners with biopharma to provide multi-omic data that is fueling the next wave of biotherapeutics. Gain actionable insights and build tailored solutions for each phase of development.
Have Questions?
†Not a replacement for comprehensive germline testing.
Incidental pathogenic alterations are reported, including ACMG recognized cancer genes.
Negative results do not imply the patient does not harbor a germline mutation.